How computing environment are usually described?
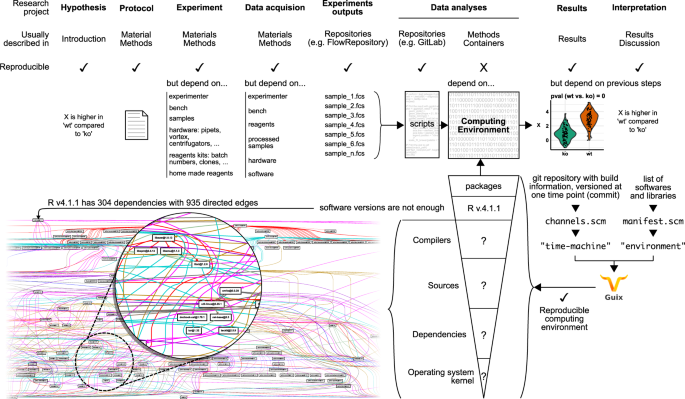
Share version labels. In most paper, software and packages used for the analyses are described with versions labels. But to be built or used, software rely on other software that also rely on other software and so on. This may seem anecdotal but, R V4.1.1 (without any packages) need at least 304 other software to be built and to run. Thus, any other user who will install the same R version will fail to obtain the exact same environment that was used in your research.
Share self-contained environment. Others provide a self-contained environment with a container image. Containers cannot be examined and do not provide information on how they were compiled. Thus, if containers are lost, it will be hard, near impossible, to rebuild them. Unless, one recovers the old computer with the same software from the time in which container was built.
Altogether, these two solutions may lead to the impossibility to reproduce results in short-term and long-term.
What is new with Guix?
Guix tackles these two issues by providing a simple practice that allows any users to share the recipe to build the same computing environment. Guix uses defined state dependencies that will indicate which versions of all dependencies were used. Then, by providing the list of software needed in your environment, Guix builds the computing environment using the same dependencies.
Is it practical and easy to use?
Guix practice is straightforward. Only two text files are required to:
- Describe the fixed state of dependencies
- Describe all software and packages that are required to build the environment
The Git repository which accompanies this article provides examples to start using Guix or to show how we reproduced previously published results using Guix.
Concrete illustration of how we used Guix in our research can be found in this work from our group: https://doi.org/10.1182/blood.2022016926
Where to start?
New users can start with the R_workflow folder from our Git repository or read the website of Guix usage in scientific context (Guix-HPC) for further information.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in